1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
| rm(list = ls()) #### 魔幻操作,一键清空~
getwd()
setwd('C:/Users/Administrator/project_gdc2')
#先把下载的临床数据文件解压
tar_file <- "clinical.cart.2024-12-22.tar.gz"# 导入tar.gz文件
dir.create('clinical_data')
extract_dir <- "./clinical_data" #解压后存放位置
untar(tar_file, exdir = extract_dir) #解压tar.gz文件
# 提取临床数据整理生存分析需要的数据
setwd("C:/Users/Administrator/project_gdc2/clinical_data") ##设置路径
library(readr)
library(dplyr)
#将下载好的metadata.json文件放入clinical文件夹
# install.packages("jsonlite")
library(jsonlite)
json <- jsonlite::fromJSON("../metadata.cart.2024-12-22.json") #读取JSON文件
entity_submitter_id <- sapply(json$associated_entities, function(x) unlist(x[, 1]))
case_id <- sapply(json$associated_entities, function(x) unlist(x[, 3]))
sample_case <- t(rbind(entity_submitter_id, case_id))
clinical <- read_tsv('clinical.tsv') #读取tsv文件
clinical <- as.data.frame(clinical[!duplicated(clinical$case_id),]) #去除重复的sample
# 371
str(sample_case) # 查看sample_case的结构
str(clinical) # 查看clinical的结构
sample_case <- as.data.frame(sample_case)
## 将sample_case$case_id和clinical$case_id转化为字符串格式,便于后续操作。
sample_case$case_id <- as.character(sample_case$case_id)
clinical$case_id <- as.character(clinical$case_id)
matrix <- merge(sample_case,clinical,by="case_id",all.x=T)
#424
colnames(clinical)
demo <- c("case_submitter_id","age_at_index","ethnicity","gender","race",
"vital_status","days_to_death","days_to_last_follow_up",
"ajcc_pathologic_stage","ajcc_pathologic_t","ajcc_pathologic_m",
"ajcc_pathologic_n","treatment_type")
matrix = matrix[,demo] #筛选需要的临床信息
dim(matrix)# 424 13
colnames(matrix) <- c("ID","Age","Ethnicity","Gender","Race",
"Status","days_to_death","days_to_last_follow_up",
"Stage","T","M","N","Treatment")
#排除结局为"Not Reported"的Sample,保留Alive和Dead的数据
matrix = matrix[matrix$Status %in% c('Alive','Dead'),]
# 422
# 把matrix数值列转换为数值型,便于记录生存信息
matrix$days_to_last_follow_up <- as.numeric(matrix$days_to_last_follow_up)
matrix$days_to_death <- as.numeric(matrix$days_to_death)
matrix$Age <- as.numeric(matrix$Age)
# 去除NA,替换为0
matrix$days_to_last_follow_up[is.na(matrix$days_to_last_follow_up)] = 0
matrix$days_to_death[is.na(matrix$days_to_death)] = 0
matrix$Age [is.na(matrix$Age )] = 0
matrix$days <- ifelse(matrix$Status=='Alive',matrix$days_to_last_follow_up,matrix$days_to_death)
## 添加生存分析需要的信息:存活状态、月、年
matrix$OS <- ifelse(matrix$Status == "Alive", 0, 1)
matrix$month=round(matrix$days/30,0) #以month为单位,小数不保留
matrix$OS.time <- floor(matrix$month/12)
floor(12.1)#12 整数年,也可以不取整。
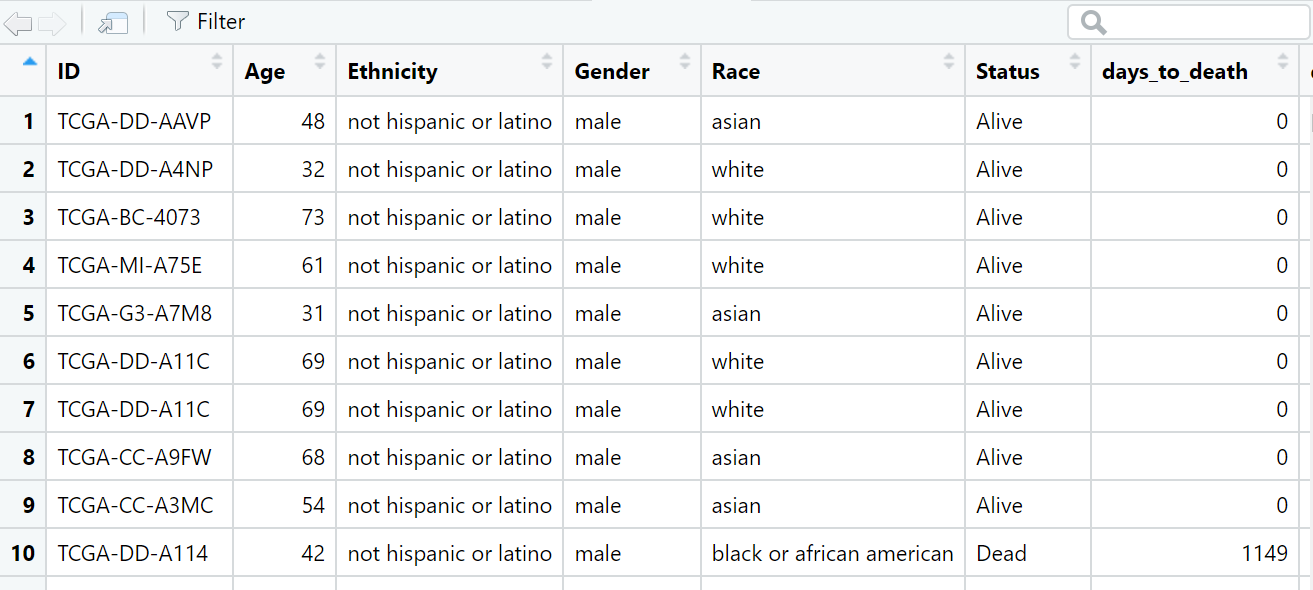
head(matrix)
# 保存临床信息
fwrite(matrix,"../RawData/03.LIHC_clin.txt") # txt格式
write.csv(matrix, "../RawData/csv/03.LIHC_clin.csv", row.names = F) # csv格式
|